Study population
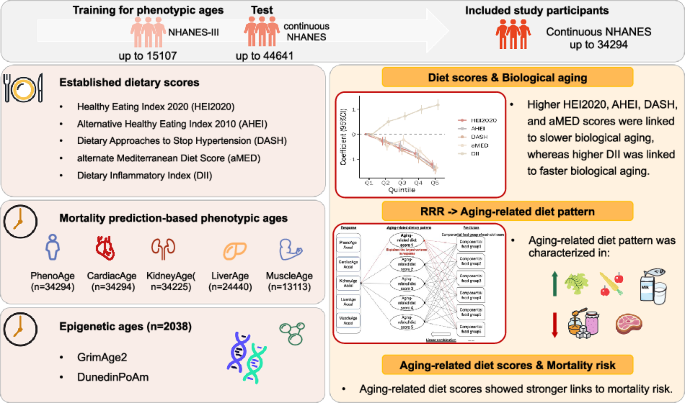
The NHANES began in the early 1960s to assess the health and nutritional status of the US population through a complex, stratified, multistage probability cluster sampling design, representing the non-institutionalized, civilian population living in the US ( Phenotypic ages (PhenoAge11,12 and organ-specific age9,10) were calculated using NHANES-III (from 1988 to 1994) as the training dataset and continuous NHANES (from 1999 to 2018) as the validation dataset (Fig. 1). An association analysis was then conducted, which included up to 34,330 participants using the continuous NHANES data. Inclusion criteria included ≥20 years old and <85 years before 2007, and <80 years from 2007 onwards. These restrictions were imposed due to age cut-offs for the NHANES coding (details seen at Additional inclusion criteria included non-pregnant, obtainable dietary scores, daily average calorie intake between 500 and 4000 kcal, available biological age biomarkers, ≥one 24-h diet recall, and exclusion of missing related covariates information from NHANES 1999–2018 (Fig. 1). The NHANES study protocol was approved by the Institutional Review Board of the National Center for Health Statistics, and all participants provided written informed consent.
Assessment of diet scores
Twenty-four-hour diet recalls were collected in the NHANES mobile examination center using the US Department of Agriculture (USDA) automated multiple-pass method24. Food groups were disaggregated using the USDA’s Food Patterns Equivalents Database and MyPyramid Equivalents Database. The USDA’s Food and Nutrient Database for Dietary Studies was used to calculate nutrient and energy intakes. Participants who completed at least one valid diet recall were included in the diet scores assessment. The first recall was used for participants with one recall, and 2-day averages were used for those with two. In this study, “Dietaryindex” was used to calculate five previously established diet scores: HEI202025, AHEI26, DASH27, aMED28, and DII29. “Dietaryindex” is a versatile and validated R package that facilitates the standardized calculation of dietary indices for use in epidemiological and clinical research (details are publicly available at https://github.com/jamesjiadazhan/dietaryindex)30. The calculation process involved two steps: (1) determining the serving sizes for each food and nutrient category; and (2) computing individual dietary indices based on these data. The HEI2020 assesses diet quality based on adherence to the US dietary guidelines 2020–2025, with 13 components scored from 0 to 100, emphasizing fruits, vegetables, whole grains, and healthy fats. The AHEI comprises 11 components, with positive scores for vegetables, fruits, whole grains, and fish, and negative scores for red meat and sugar. Scores range from 0 to 110. The DASH score is based on eight components, with positive scores for fruits, vegetables, and low-fat dairy and negative scores for sodium and red meat. Scores range from 8 to 40. The aMED is a median-based score based on nine components, with positive scores for vegetables, fruits, nuts, fish, and moderate alcohol. Scores range from 0 to 9. The DII estimates a diet’s inflammatory potential according to six inflammatory biomarkers: IL-1β, IL-4, IL-6, IL-10, TNF-α, and CRP, including micronutrients, macronutrients, and also commonly consumed bioactive components such as flavonoids and tea in standardized Z-score based on the world average and standard deviation with weighted coefficients ranging from −4.94 to 5.16, with higher values indicate more pro-inflammatory diets. Details for the calculation of all diet scores are included in the Supplementary Table 15. We also considered the planetary health diet during 2005–201830. Diet scores were evaluated both as categorical (quintiles) and continuous exposures per interquintile range (defined as the range between the 90th and 10th percentiles).
Assessment of covariates
During the survey, each participant completed a household interview and underwent a physical examination of health and nutritional status. Data on age, sex, ethnicity, education, poverty-income ratio (PIR), smoking status, and medical conditions (including hypertension, CVD, diabetes, and cancer) were collected from household interviews via structured questionnaires. Body weight, height, and blood pressure were obtained at the mobile examination center by trained staff. Body mass index was calculated as weight in kilograms divided by the square of height in meters. Blood samples were collected at the visit, and a complete blood count and biochemical analysis were performed according to the NHANES Laboratory/Medical Technologists Procedures Manual. Habitual physical activity was defined as engaging in ≥150 min of leisure activity per week. CVD and cancer were identified based on participants’ self-reports of having been informed of these conditions by a healthcare professional. Hypertension was determined by self-reported physician diagnosis, use of antihypertensive agents, or a systolic blood pressure ≥140 mmHg and/or diastolic blood pressure ≥90 mmHg at the time of visit. Diabetes was determined by self-reported physician diagnosis, use of antidiabetic agents, or glycated hemoglobin A1c ≥ 6.5% or fasting blood plasma ≥126 mg/dL.
Construction of the biological age models
In this study, we employed several established biological aging measures to capture different dimensions of aging. PhenoAge, derived from clinical biomarkers, reflects systemic mortality risk. Building on this framework, we further extended the approach to develop organ-specific ages, which assess aging at the level of individual physiological systems. Two DNA methylation-based measures were also included: GrimAge2, optimized for mortality and morbidity prediction, and DunedinPoAm, which quantifies the rate of physiological decline. Detailed procedures for the calculation of each measure are described below.
Phenotypic ages, including PhenoAge and organ-specific ages (referring to CardiacAge, KidneyAge, LiverAge, and MuscleAge in this study) according to mortality prediction-based models, are based on clinical biomarkers (both physical and physiological measures) used to reflect the health and functionality from NHANES 1999–2018. The following formula was used to: (1) Construct the Gompertz cumulative distribution function (CDF) to estimate the 10-year all-cause mortality risk (t = 10 here), with age as the predictor.
$$\mathrm{CDF}(t,\,\mathrm{age})=1-\exp (-\exp (\mathrm{age}* {{\rm{\beta }}}_{0}+{c}_{0}){{{\rm{\gamma }}}_{0}}^{-1}(\exp ({{\rm{\gamma }}}_{0}t)-1))$$
(1)
(2) Construct the Gompertz CDF to estimate the 10-year all-cause mortality risk (t = 10 here), with CA (chronological age), and biomarkers as the predictors. xb was designated the linear combination of CA and other biomarkers.
$$\mathrm{CDF}(t,\,\mathrm{xb})=1-\exp (-\exp (\mathrm{xb})* {{{\rm{\gamma }}}_{1}}^{-1}* (\exp ({{\rm{\gamma }}}_{1}t)-1))$$
(2)
$$\mathrm{xb}={\beta }_{1}* \mathrm{CA}+{\beta }_{2}* \mathrm{biomarkers}+{c}_{{1}}$$
(3) By setting ① = ②, the Gompertz CDF for age was equated with that for CA and biomarkers at 10 years. Through a series of transformations, the explicit expression was derived for the predicted age as follows:
$$\mathrm{predicted}\,\mathrm{age}=\frac{1}{{{\rm{\beta }}}_{{0}}}\mathrm{ln}\frac{{{{\rm{\gamma }}}_{1}}^{-1}(\exp ({{\rm{\gamma }}}_{1}t)-1)}{{{{\rm{\gamma }}}_{0}}^{-1}(\exp ({{\rm{\gamma }}}_{0}t)-1)}-\frac{{c}_{0}}{{{\rm{\beta }}}_{0}}+\frac{\mathrm{xb}}{{{\rm{\beta }}}_{0}}$$
(3)
Following the approach, a mortality prediction-based model was trained in the NHANES-III using systemic/organ-specific biomarkers, and applied it in the continuous NHANES. A predicted age in the model represents the chronological age in the reference population that corresponds to his/her 10-year mortality risk, as determined by systemic/organ-specific biomarkers. Based on previously identified data, the following systemic/organ-specific biomarkers9,10 were used: (i) Systemic biomarkers for PhenoAge: albumin, creatinine, serum glucose, log-transformed CRP (due to high missingness, PhenoAge was only considered with CRP in the sensitivity analysis), lymphocyte percentage, mean red cell volume, red cell distribution width, alkaline phosphatase, and white blood cell count. (ii) Cardiovascular biomarkers for CardiacAge: systolic blood pressure, diastolic blood pressure, total cholesterol, high-density lipoprotein cholesterol, and pulse rate. (iii) Kidney biomarkers for KidneyAge: uric acid, serum albumin, serum creatinine, and blood urea nitrogen. (iv) Liver biomarkers for LiverAge: aspartate transaminase, alanine transaminase, gamma-glutamyl transferase, serum albumin, total bilirubin, and alkaline phosphatase. (v) Musculoskeletal biomarkers for MuscleAge: body weight, body mass index, waist circumference, waist-to-hip ratio, femur bone mineral density, alkaline phosphatase, and serum calcium. Additionally, constructed were several commonly used first-generation models (elastic net, random forest, and XGBoost) with chronological age as a training label and compared with the mortality prediction-based model (Supplementary Fig. 5) and cross-sectional associations with chronic diseases (Supplementary Fig. 6). These three methods were chosen because they are widely applied in aging-clock research and are representative of distinct methodological families: penalized linear regression (elastic net), bagging-based tree ensembles (random forest), and XGBoost31,32. To enable comparison between methods, age accelerations were standardized by dividing by the standard deviation.
DNA methylation (DNAm) data from 1999 to 2002 ( were used to investigate potential mechanisms or indicators, which included GrimAge233 and DunedinPoAm34. DNAm was measured using Illumina EPIC BeadChip arrays for a subgroup of adults ≥50 years, and surveyed in 1999–2000 or 2001–2002, whose blood samples were eligible for DNA isolation. Briefly, GrimAge2 was a linear equation of chronological age, sex, and ten DNAm biomarkers, including ten DNAm-based surrogates for smoking pack-years, adrenomedullin levels, beta-2 microglobulin, cystatin C, growth differentiation factor 15, leptin, log-scale high-sensitivity CRP, log-scale hemoglobin A1C, plasminogen activation inhibitor 1, and tissue inhibitor metalloproteinase 1. DunedinPoAm employed methylation sites to quantify the pace of aging, through an analysis of 18 longitudinal biomarkers including glycated hemoglobin, cardiorespiratory fitness, waist-hip ratio, FEV1/FVC ratio, FEV1, mean arterial pressure, body mass index, leukocyte telomere length, creatinine clearance, blood urea nitrogen, lipoprotein (a), triglycerides, gum health, total cholesterol, white blood cell count, high-sensitivity CRP, HDL cholesterol, ApoB100/ApoA1 ratio.
Biological age acceleration was calculated as the residuals from linear regressions of predicted biological age on chronological age, stratified by sex. A higher age acceleration, compared to peers of the same sex and chronological age, indicated accelerated aging.
Ascertainment of mortality
The survival data were extracted from the National Center for Health Statistics Data 2019 Public-Use Linked Mortality Files ( For each participant, person-time was calculated from the date of the baseline survey interview to the date of death or the end of follow-up (December 31, 2019), whichever occurred first.
Statistical analysis
The appropriate sample weights, stratification, and clustering of the complex survey design were incorporated in all analyses to ensure national estimates. The family PIR was missing in >5% of the participants, and was assigned an independent category. All other missing covariates were coded with median values for continuous variables or mode values for categorical variables. Demographic characteristics were analyzed across quintiles (top and bottom) of diet scores. Data were presented as weighted means (SE) for continuous variables and unweighted sample sizes (weighted percentage) for categorical variables. The weighted Pearson’s correlation (weighted ρ) was used to test the correlations within dietary scores or within phenotypic age accelerations.
Weighted multivariable linear regression models were used to examine the associations of five previously established diet scores with accelerations of PhenoAge and organ-specific age (1999–2018) and epigenetic measures (1999–2002) in the continuous NHANES. Two models were constructed to account for potential confounding: model 1 was adjusted for age (continuous) and sex (women; men) and model 2 was further adjusted for ethnicity (non-Hispanic White; non-Hispanic Black; Mexican American; other Hispanic; others), education (beyond high school; up to high school), PIR (<1.00; ≥1.00), smoking status (never; former; current), physical activity (regular exercise; or not), body mass index (<30 kg/m2; ≥30 kg/m2), hypertension (yes; no), CVD (yes; no), diabetes (yes; no), cancer (yes; no), and average caloric intake (continuous) based on model 1. Linear trends were tested by treating the median value of each diet score category as a continuous variable.
RRR was used to identify aging-related diet scores using componential food groups within each of five previously established diet scores after standardizing for both the predictors (food groups) and response (accelerations of phenotypic ages at systemic and organ-specific levels) (Supplementary Fig. 10). Factor loadings, representing the standardized correlations between component groups and dietary patterns (factors), were computed. Selected was the factor that explained the greatest variation in the aging pattern. Additionally, the proportion of variance explained by the selected factor for both the response variables and component groups was calculated. Aging-related diet scores were derived from the linear combination of predictors and corresponding loading coefficients. Food/nutritional groups with absolute loading values greater than 0.2 were considered important for identifying aging-related diet score35.
In the prospective study, HRs and 95% CIs for the associations between diet quality and mortality were calculated using weighted Cox proportional hazards regression models, adjusting for the same covariates as mentioned above. The proportional hazards assumption was assessed using Schoenfeld tests of interaction between each variable with time, which revealed no violations. Stratified analyses were conducted to assess potential effect modification by the covariates, with P for interaction estimated using the likelihood ratio (Rao-Scott) test by adding a product term between the stratifying variable and diet score in the models. In the sensitivity analysis, the sample was further restricted to participants whose food consumption on the previous day was typical of their usual diet, to minimize random measurement errors from self-reported diet assessments. Also conducted was a mediation analysis to explore whether and how much epigenetic age acceleration mediates the association between diet score and mortality risk using the R “CMAverse” package. The method estimated effects on the HRs and 95% CIs, with two models: a mediator model (linear regression) and an outcome model (Cox regression). The mediator model examined the relationship between the exposure (diet scores, continuous) and mediators (epigenetic age acceleration, continuous), adjusting for the covariates in the full model, while the outcome model examined the relationship between the mediator and mortality risk (survival data), with the exposure, the mediator, and the covariates in the full model adjusted. The total effect was decomposed into direct effect (DE) and indirect effect (IDE), with IDE representing the effect of individual diet score on mortality risk that was explained by the mediators in the model, while DE represented the diet score’s effect on mortality risk independent of the mediator. Nonparametric bootstrapping (500 times) was used to estimate 95% CI and P values. The method quantified the extent of mediation by calculating the proportion of the association attributable to the mediator (DE × (IE−1)/[TE−1]).
All the analyses were conducted in R version 4.1.3. A two-sided P value < 0.05 was considered statistically significant.
link